Characterization of Arabidopsis thaliana promoter bidirectionality and antisense RNAs by depletion of nuclear RNA decay enzymes | bioRxiv

Integration of Single-Cell RNA- and CAGE-seq Reveals Tooth-Enriched Genes - Y. Chiba, K. Yoshizaki, T. Tian, K. Miyazaki, D. Martin, , Genomics and Computational Biology Core, Genomics and Computational Biology Core, K.
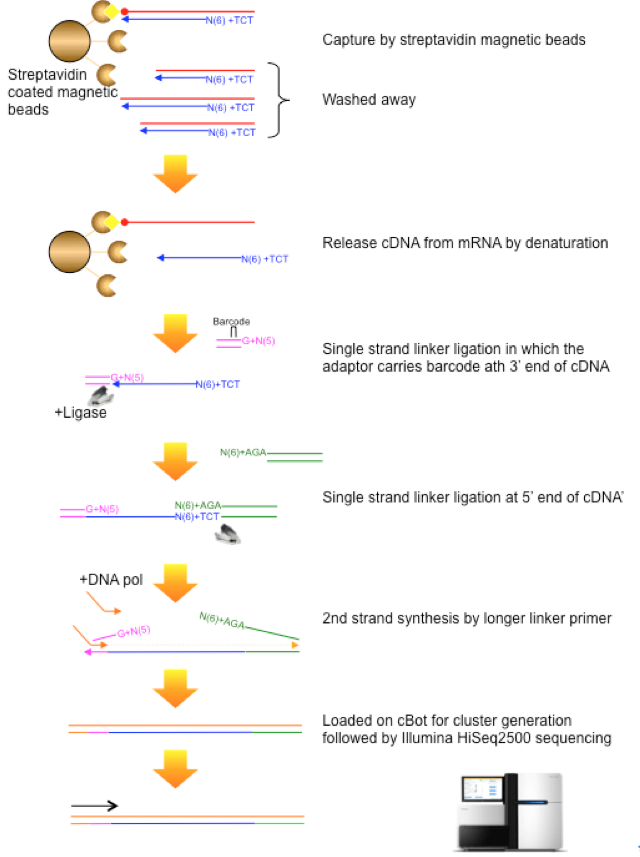
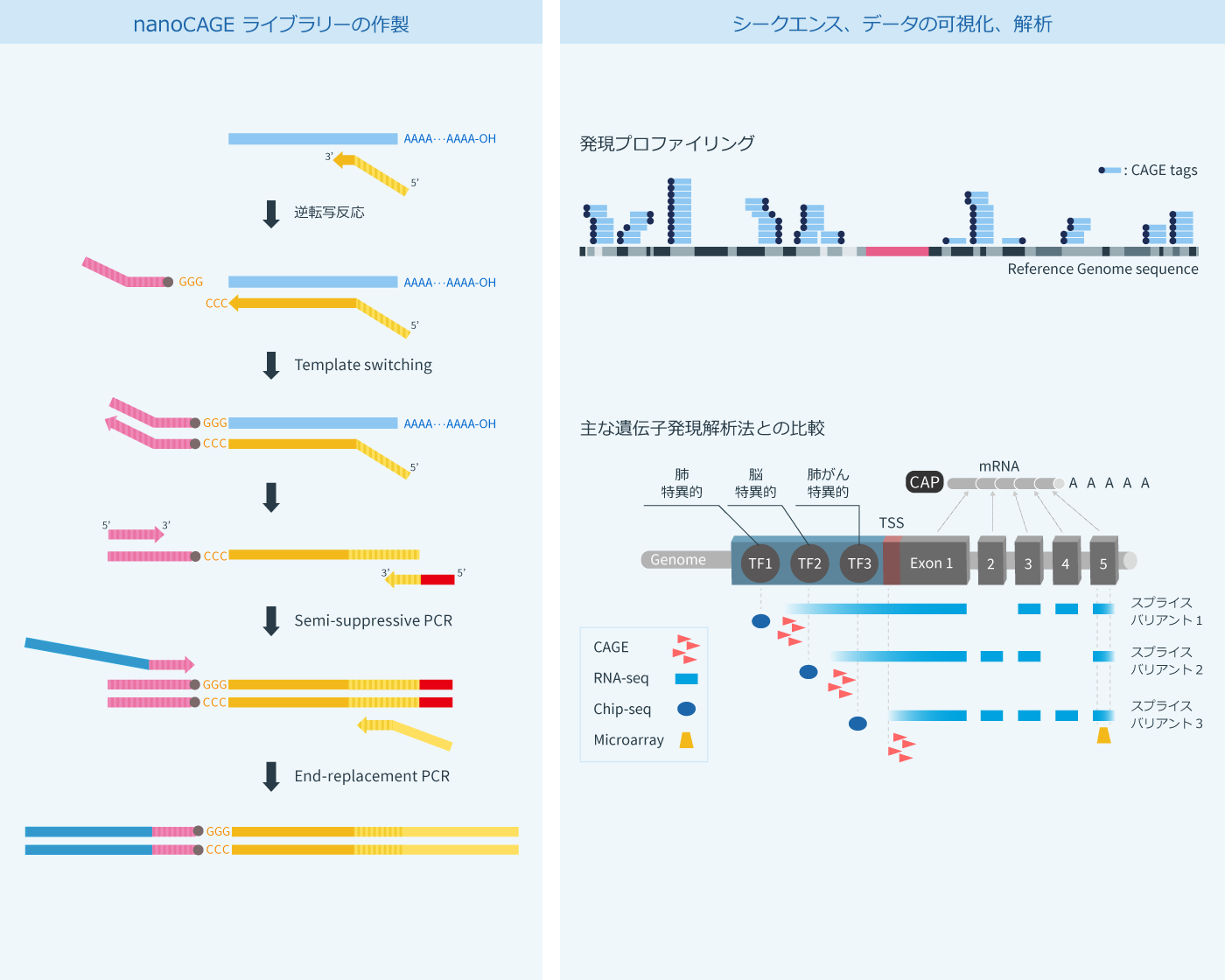
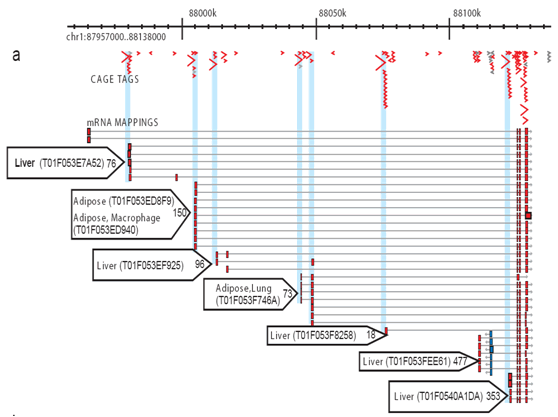
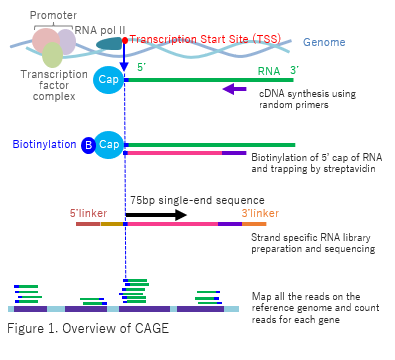
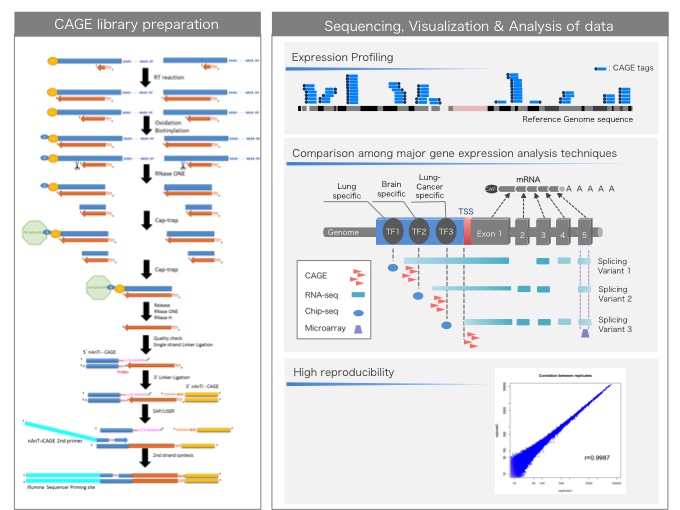
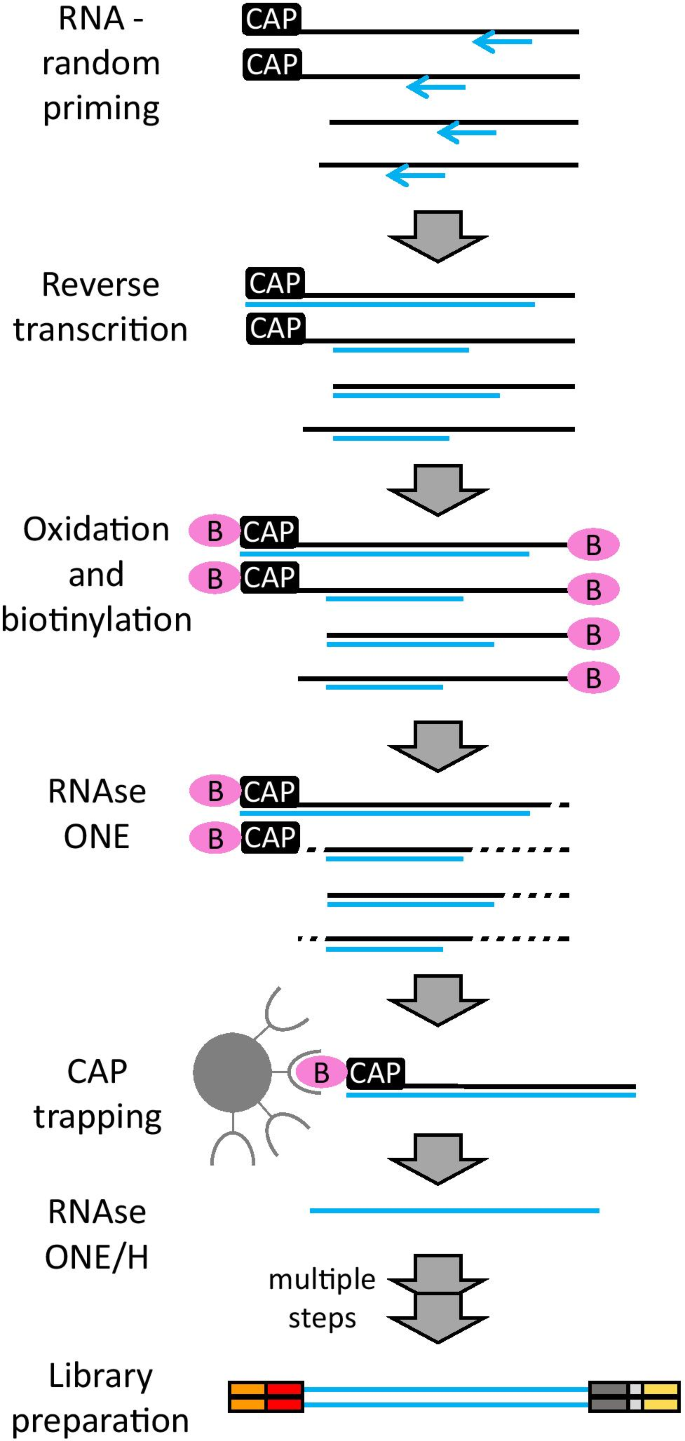
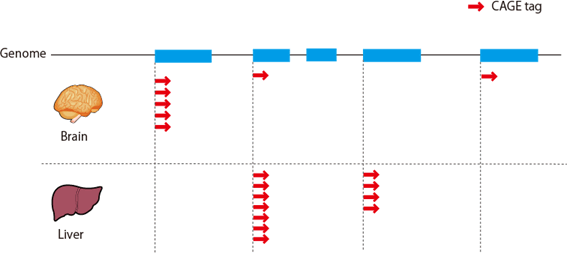
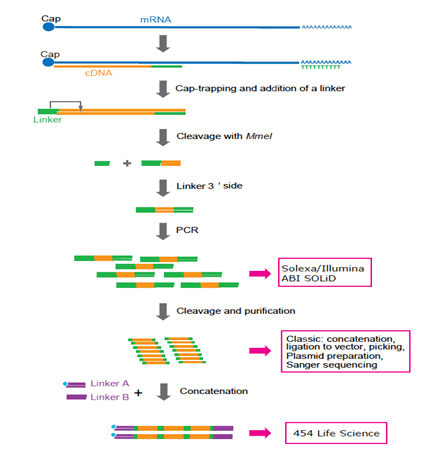
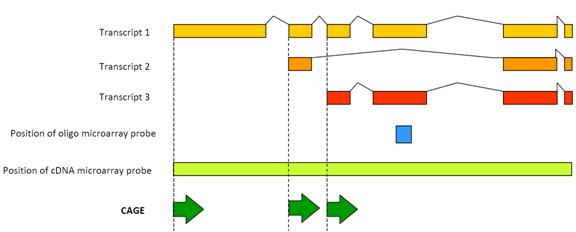
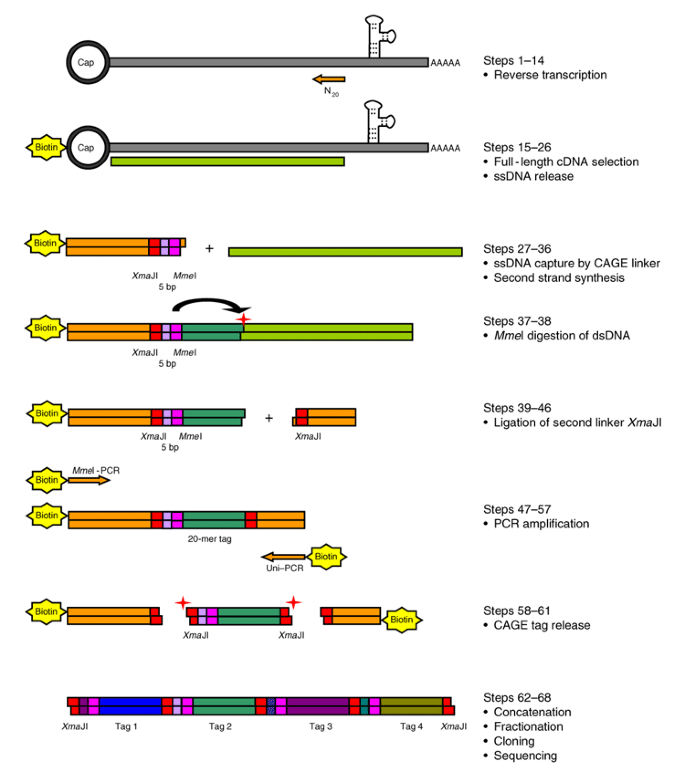
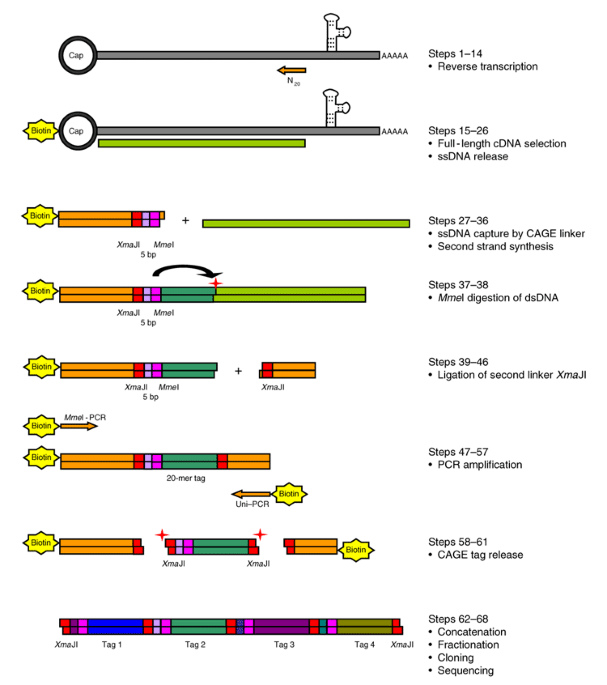
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター

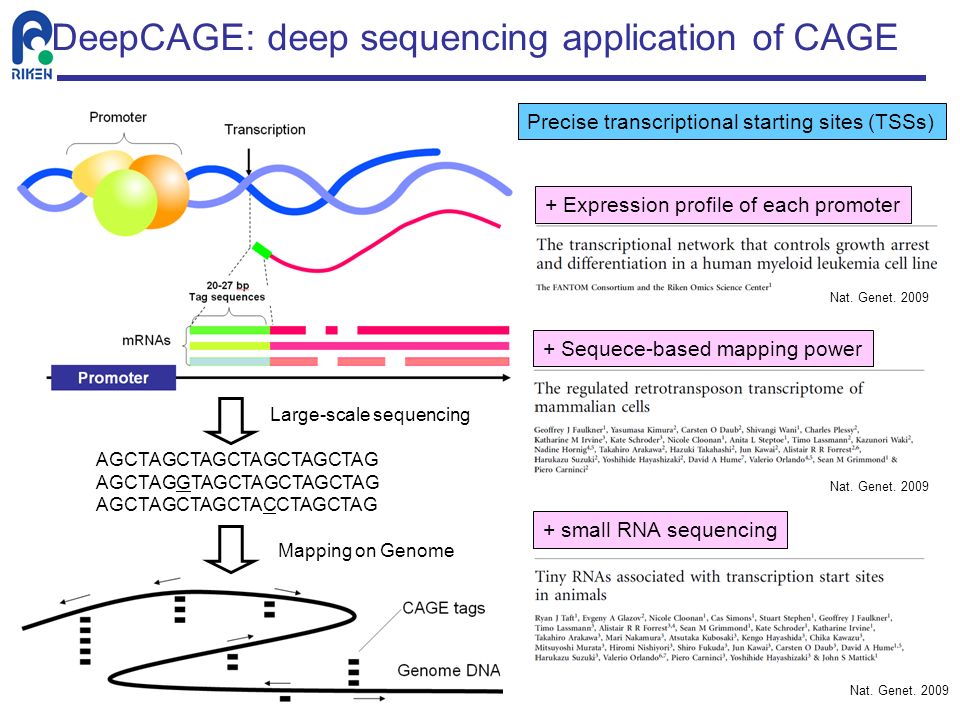
DeepCAGE: Incorporating Transcription Factors in Genome-wide Prediction of Chromatin Accessibility | bioRxiv

DeepCAGE: Incorporating Transcription Factors in Genome-wide Prediction of Chromatin Accessibility - ScienceDirect
![PDF] piPipes: a set of pipelines for piRNA and transposon analysis via small RNA-seq, RNA-seq, degradome- and CAGE-seq, ChIP-seq and genomic DNA sequencing | Semantic Scholar PDF] piPipes: a set of pipelines for piRNA and transposon analysis via small RNA-seq, RNA-seq, degradome- and CAGE-seq, ChIP-seq and genomic DNA sequencing | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/ae869ecce6a4a05d8ac4db3ebc27bb833beef92e/2-Figure1-1.png)
PDF] piPipes: a set of pipelines for piRNA and transposon analysis via small RNA-seq, RNA-seq, degradome- and CAGE-seq, ChIP-seq and genomic DNA sequencing | Semantic Scholar